Molecular and cell biology of acid (SMPD-1) and neutral sphingomyelinases (SMPD-2 and SMPD-3)
The phosphosphingolipid class of sphingomyelins (SM) resembles abundant structural element of all eukaryotic membrane lipid bilayers. SMs are cleaved in phospholipase c like reaction to ceramides and phosphorylcholin. Ceramides regarded as lipid signalling molecules in developmental and apoptotic pathways.
To understand the enigmatic role of SM and of acid and neutral sphingomyelinases in cellular metabolism and particularly with respect to their proposed role as key players in apoptotic signalling pathways we generated the acid sphingomyelin mouse model (smpd1-/-), the phenotype of which revealed a mimicry of the lysosomal storage disease, known as human Niemann-Pick disease, type A.
We further discovered and cloned the ubiquitously occurring neutral sphingomyelinase 1 (nSmase1) (smpd-2) and nSmase2 (smpd-3). We subsequently generated the respective null allelic mouse mutant models smpd2-/- and smpd3-/-. Crossing the two genotypes yielded the smpd2-/-smpd3-/- double mutant, which proved to be free of Mg++ dependent neutral sphingomyelinase activity.
The smpd3-/- and smpd2-/-smpd3-/- double mutants develop general postnatal dwarfism, hypoplasia and growth retardation, no altered apoptosis was observed. We have identified SMPD3 expression in hypothalamic neurosecretory neurons as a novel check point in the hypothalamus pituitary growth axis.
SMPD3, a novel essential player in Golgi vesicle transport
Comprehensive phenotyping data unveiled SMPD3 as regulating element in Golgi vesicle transport with peptide hormones (growth hormone, GHRH- corticotropin-CRH, gonadotropin- GnRH, thyrotropin releasing hormone, TRH) as cargo in hypothalamic neuro-secretory neurons.
Importantly, chondrocytes autonomously express SMPD3 in Golgi stacks, critically involved in vesicular secretion of extracellular matrix proteins essential for the longitudinal extension in epiphyseal growth plate.
Different from the lysosomal "house keeping" sphingomyelinase, SPMD3 enzyme activity is regulated. Smpd3 specific cDNA under the control of the chondrocyte specific promoter Coll2A normalized the phenotype in the smpd3-/- mouse. This underlines the important role of cell specific and developmental regulation of smpd3 expression.
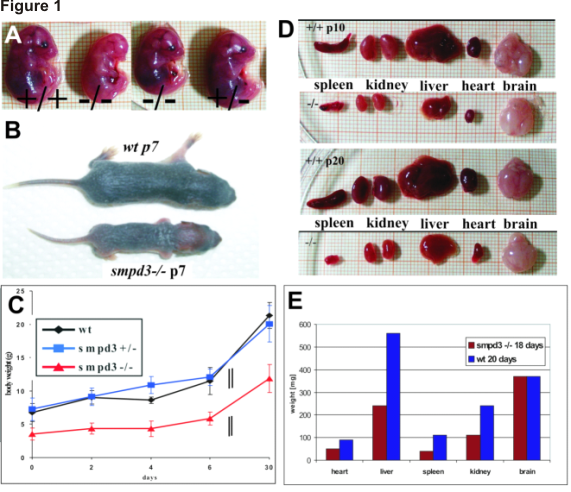
Figure 1
(A) Comparison of e14 embryos smpd3+/+, smpd3-/- and smpd3+/- siblings. (B) Severe postnatal growth retardation of p7 littermates. (C) Growth curves of wt, smpd3+/- and smpd3-/- male and female littermates. (D) Hypoplasia of spleen, kidney, liver heart and brain of +/+ and -/- p10 and p20 mice. (E) Weight of organs of +/+ and -/- mice. (F) Serum IGF1 concentrations of wt and smpd3-/- male siblings. (G) cell count of splenocytes at 1, 6 and 24 month of age and (H) cell count of pituicytes at 1 and 24 month of age.
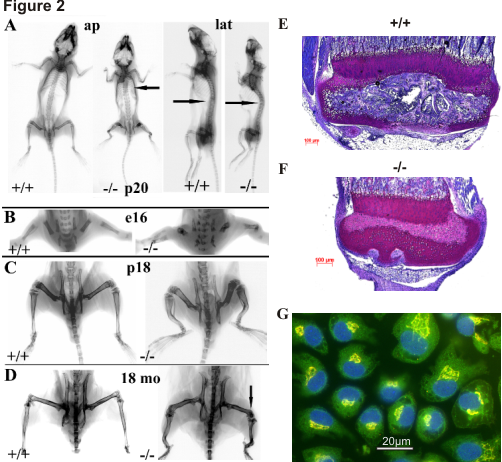
Figure 2
Short-limbed dwarfism and severe skeletal deformation in smpd3-/- mice (A-D). (A) Anteroposterior X-ray imaging reveals rhizomelic long bones and a cage-shaped thorax (arrow) in p20 smpd3-/- mice. (B) Skeletal phenotype observed during late embryonic development (e16). (C) Short stature and deformation of femora, tibiae and humeri in juvenile (p18) and (D) adult (18mo) mice. (E) Proximal tibia of smpd3+/+ and (F) smpd3-/- mice. (G) IHC of SMPD3 subcellular localization. The SMPD3 is strictly located in the Golgi stacks.
Literature
W. Stoffel, B. Jenke, B. Holz, E. Binczek, R. Gunter, J. Knifka, J. Koebke, A. Niehoff (2007) Neutral sphingomyelinase (SMPD3) deficiency causes a novel form of chondrodysplasia and dwarfism that is rescued by Col2A1-driven smpd3 transgene expression Am J Pathol. 171,153-61
W. Stoffel, B. Jenke, B. Blöck, M. Zumbansen, J. Koebke (2005) Neutral sphingomyelinase 2 (smpd3) in the control of postnatal growth and development. Proc. Natl. Acad. Sci. USA 120, 4554-4559
M. Zumbansen, W. Stoffel (2002) Neutral sphingomyelinase 1 deficiency in the mouse causes no lipid storage disease Mol. Cell Biol. 22, 3633-3638
M. Nix, W. Stoffel (2000) Perturbation of membrane microdomains reduces mitogenic signaling and increases susceptibility to apoptosis after T cell receptor stimulation Cell Death and Differentiation 7, 413-424
S. Tomiuk, M. Zumbansen, W. Stoffel (2000) Characterization and subcellular localization of murine and human magnesium-dependent neutral sphingomyelinase J. Biol. Chemistry 275, 5710-5717
K. Hofmann, S. Tomiuk, G. Wolff, W. Stoffel (2000) Cloning and characterization of the mammalian brain-specific, Mg2+-dependent neutral sphingomyelinase Proc. Natl. Acad. Sci. USA 97, 5895-5900
B. Stoffel, P. Bauer, M. Nix, K. Deres, W. Stoffel (1998) Ceramide-independent CD28 and TCR signaling but reduced IL-2 secretion in T cells of acid sphingomyelinase-deficient mice Eur. J. Immunol. 28, 874-880
S. Tomiuk, K. Hofmann, M. Nix, M. Zumbansen, W. Stoffel (1998) Cloned mammalian neutral sphingomyelinase: Functions in sphingolipid signaling? Proc.Natl.Acad. Sci.USA 95, 3638-3643
T. Kümmel, J. Thiele, R. Schroeder, W. Stoffel (1997) Pathology of visceral organs and bone marrow in an acid sphingomyelinase deficient knock-out mouse line, mimicking human Niemann-Pick Disease type a. A light and electron microscopic study Pathology Research and Practice 193, 663-671
M. Zumbansen, W. Stoffel (1997) Tumor necrosis factor a activates NF-?B in acid sphingomyelinase-deficient mouse embryonic fibroblasts J.Biol.Chem. 272, 10904-10909
Th. Kümmel, R. Schroeder, W. Stoffel (1997) Light and electron microscopic analysis of the central and peripheral nervous systems of acid sphingomyelinase-deficient mice resulting from gene targeting Journal of Neuropathology and Experimental Neurology 56, 171-179
D. Newrzella, W. Stoffel (1996) Functional analysis of the glycosylation of murine acid sphingomyelinase J. Biol.Chem. 271, 32089-32095
B. Otterbach, W. Stoffel (1995) Acid sphingomyelinase-deficient mice mimic the neurovisceral form of human lysosomal storage disease (Niemann-Pick disease) Cell 81, 1053-1061
D. Newrzella, W. Stoffel (1992) Molecular cloning of the acid sphingomyelinase of the mouse and the organization and complete nucleotide sequence of the gene. Biol. Chem. Hoppe-Seyler, 373, 1233-1238